The Database Class¶
A Class designed for the storing, retrieval and updating of results.
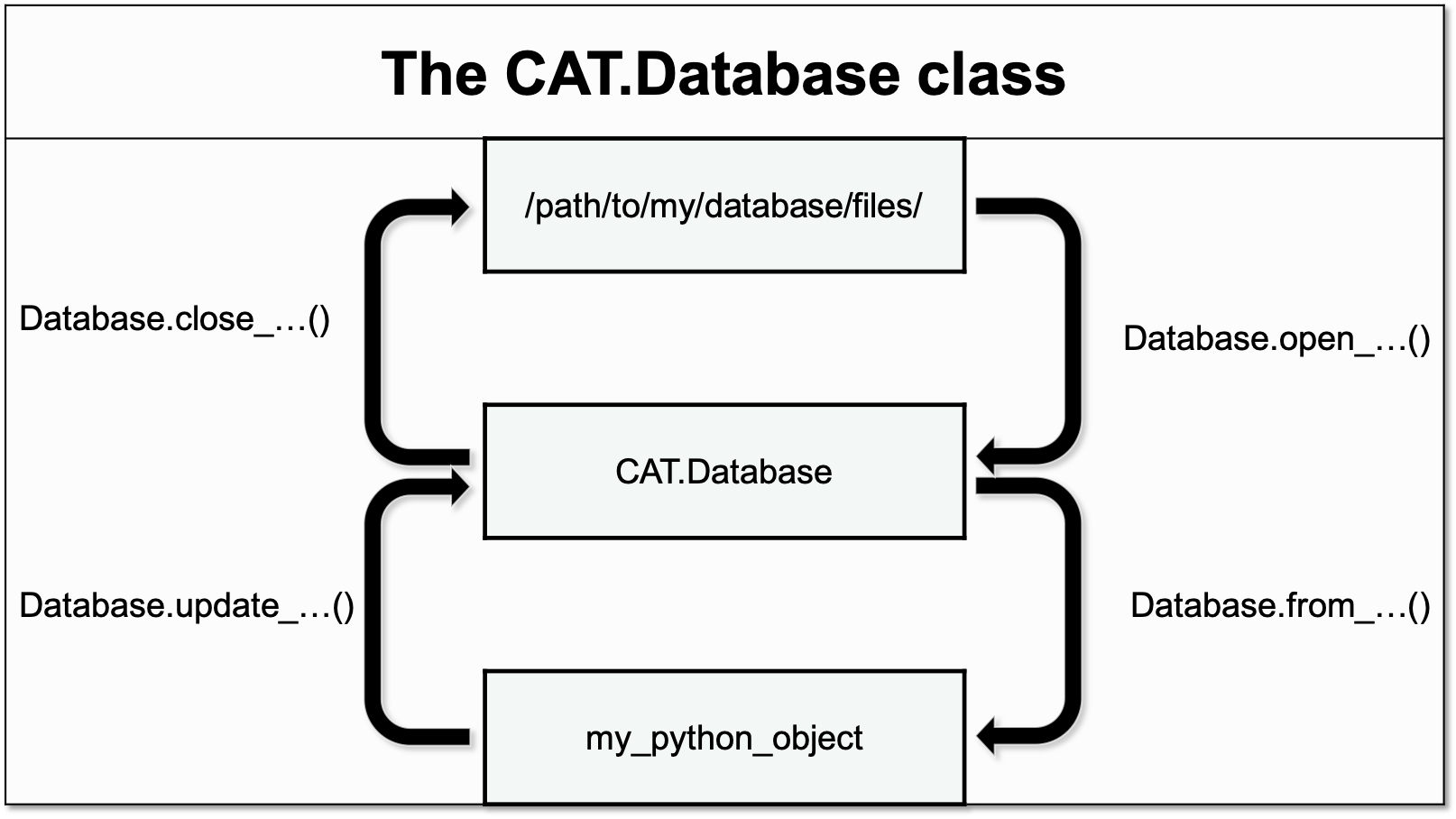
The methods of the Database class can be divided into three categories accoring to their functionality:
Opening & closing the database - these methods serve as context managers for loading and unloading parts of the database from the harddrive.
The context managers can be accessed via the
MetaManager.open()method ofDatabase.csv_lig,Database.csv_qd,Database.yamlorDatabase.hdf5, with the option of passing additional positional or keyword arguments.>>> import CAT >>> database = CAT.Database() >>> with database.csv_lig.open(write=False) as db: >>> print(repr(db)) DFCollection(df=<pandas.core.frame.DataFrame at 0x7ff8e958ce80>) >>> with database.yaml.open() as db: >>> print(type(db)) <class 'scm.plams.core.settings.Settings'> >>> with database.hdf5.open('r') as db: >>> print(type(db)) <class 'h5py._hl.files.File'>
Importing to the database - these methods handle the importing of new data from python objects to the Database class:
update_csv()update_yaml()update_hdf5()update_mongodb()Exporting from the database - these methods handle the exporting of data from the Database class to other python objects or remote locations:
from_csv()from_hdf5()
Index¶
dirname |
|
csv_lig |
|
csv_qd |
|
hdf5 |
|
yaml |
|
mongodb |
|
update_mongodb([database, overwrite]) |
Export ligand or qd results to the MongoDB database. |
update_csv(df[, database, columns, …]) |
Update Database.csv_lig or Database.csv_qd with new settings. |
update_yaml(job_recipe) |
Update Database.yaml with (potentially) new user provided settings. |
update_hdf5(df[, database, overwrite, opt]) |
Export molecules (see the "mol" column in df) to the structure database. |
from_csv(df[, database, get_mol, inplace]) |
Pull results from Database.csv_lig or Database.csv_qd. |
from_hdf5(index[, database, rdmol]) |
Import structures from the hdf5 database as RDKit or PLAMS molecules. |
df_collection.get_df_collection(df) |
Return a mutable collection for holding dataframes. |
database_functions.as_pdb_array(mol_list[, …]) |
Convert a list of PLAMS molecule into an array of (partially) de-serialized .pdb files. |
database_functions.from_pdb_array(array[, rdmol]) |
Convert an array with a (partially) de-serialized .pdb file into a molecule. |
database_functions.sanitize_yaml_settings(…) |
Remove a predetermined set of unwanted keys and values from a settings object. |
Class API¶
Database¶
-
class
dataCAT.database.Database(path=None, host='localhost', port=27017, **kwargs)[source]¶ The Database class.
Parameters: - path (str) – The path+directory name of the directory which is to contain all database components
(see
Database.dirname). - host (str) – Hostname or IP address or Unix domain socket path of a single mongod or
mongos instance to connect to, or a mongodb URI, or a list of hostnames mongodb URIs.
If host is an IPv6 literal it must be enclosed in
"["and"]"characters following the RFC2732 URL syntax (e.g."[::1]"for localhost). Multihomed and round robin DNS addresses are not supported. SeeDatabase.mongodb. - port (str) – port number on which to connect.
See
Database.mongodb. - **kwargs –
Optional keyword argument for pymongo.MongoClient. See
Database.mongodb.
-
csv_lig¶ A dataclass for accesing the context manager for opening the .csv file containing all ligand related results.
Type: dataCAT.MetaManager
-
csv_qd¶ A dataclass for accesing the context manager for opening the .csv file containing all quantum dot related results.
Type: dataCAT.MetaManager
-
yaml¶ A dataclass for accesing the context manager for opening the .yaml file containing all job settings.
Type: dataCAT.MetaManager
-
hdf5¶ A dataclass for accesing the context manager for opening the .hdf5 file containing all structures (as partiallize de-serialized .pdb files).
Type: dataCAT.MetaManager
-
mongodb¶ Optional: A dictionary with keyword arguments for pymongo.MongoClient. Defaults to
Noneif aServerSelectionTimeoutErroris raised when failing to contact the host. See the host, port and kwargs parameter.Type: dict
-
update_mongodb(database='ligand', overwrite=False)[source]¶ Export ligand or qd results to the MongoDB database.
Examples
>>> from CAT import Database >>> db = Database(**kwargs) # Update from db.csv_lig >>> db.update_mongodb('ligand') # Update from a lig_df, a user-provided DataFrame >>> db.update_mongodb({'ligand': lig_df}) >>> print(type(lig_df)) <class 'pandas.core.frame.DataFrame'>
Parameters: - database (str or dict [str, pd.DataFrame]) – The type of database.
Accepted values are
"ligand"and"QD", openingDatabase.csv_ligandDatabase.csv_qd, respectivelly. Alternativelly, a dictionary with the database name and a matching DataFrame can be passed directly. - overwrite (bool) – Whether or not previous entries can be overwritten or not.
Return type: None- database (str or dict [str, pd.DataFrame]) – The type of database.
Accepted values are
-
update_csv(df, database='ligand', columns=None, overwrite=False, job_recipe=None, opt=False)[source]¶ Update
Database.csv_ligorDatabase.csv_qdwith new settings.Parameters: - df (pd.DataFrame) – A dataframe of new (potential) database entries.
- database (str) – The type of database; accepted values are
"ligand"(Database.csv_lig) and"QD"(Database.csv_qd). - columns (Sequence) – Optional: A list of column keys in df which
(potentially) are to be added to this instance.
If
None: Add all columns. - overwrite (bool) – Whether or not previous entries can be overwritten or not.
- job_recipe (plams.Settings) – Optional: A
Settingsinstance with settings specific to a job. - opt (bool) – WiP.
Return type: None
-
update_yaml(job_recipe)[source]¶ Update
Database.yamlwith (potentially) new user provided settings.Parameters: job_recipe (plams.Settings) – A settings object with one or more settings specific to a job. Returns: A dictionary with the column names as keys and the key for Database.yamlas matching values.Return type: dict_
-
update_hdf5(df, database='ligand', overwrite=False, opt=False)[source]¶ Export molecules (see the
"mol"column in df) to the structure database.Returns a series with the
Database.hdf5indices of all new entries.Parameters: - df (pd.DataFrame) – A dataframe of new (potential) database entries.
- database (str) – The type of database; accepted values are
"ligand"and"QD". - overwrite (bool) – Whether or not previous entries can be overwritten or not.
Returns: A series with the indices of all new molecules in
Database.hdf5.Return type: pd.Series_
-
from_csv(df, database='ligand', get_mol=True, inplace=True)[source]¶ Pull results from
Database.csv_ligorDatabase.csv_qd.Performs in inplace update of df if inplace =
True, thus returingNone.Parameters: - df (pd.DataFrame) – A dataframe of new (potential) database entries.
- database (str) – The type of database; accepted values are
"ligand"and"QD". - get_mol (bool) – Attempt to pull preexisting molecules from the database. See the inplace argument for more details.
- inplace (bool) – If
Trueperform an inplace update of the"mol"column in df. Otherwise return a new series of PLAMS molecules.
Returns: Optional: A Series of PLAMS molecules if get_mol =
Trueand inplace =False.Return type:
-
from_hdf5(index, database='ligand', rdmol=True)[source]¶ Import structures from the hdf5 database as RDKit or PLAMS molecules.
Parameters: - index (list [int]) – The indices of the to be retrieved structures.
- database (str) – The type of database; accepted values are
"ligand"and"QD". - rdmol (bool) – If
True, return an RDKit molecule instead of a PLAMS molecule. - close (bool) – If the database component (
Database.hdf5) should be closed afterwards.
Returns: A list of PLAMS or RDKit molecules.
Return type:
-
hdf5_availability(timeout=5.0, max_attempts=None)[source]¶ Check if a .hdf5 file is opened by another process; return once it is not.
If two processes attempt to simultaneously open a single hdf5 file then h5py will raise an
OSError.The purpose of this method is ensure that a .hdf5 file is actually closed, thus allowing the
Database.from_hdf5()method to safely access filename without the risk of raising anOSError.Parameters: Raises: OSError – Raised if max_attempts is exceded.
Return type: None
- path (str) – The path+directory name of the directory which is to contain all database components
(see
DFCollection¶
-
class
dataCAT.df_collection._DFCollection(df)[source]¶ A mutable collection for holding dataframes.
Parameters: df (pd.DataFrame) – A Pandas DataFrame (see _DFCollection.df).-
df¶ A Pandas DataFrame.
Type: pd.DataFrame
Warning
The
_DFCollectionclass should never be directly called on its own. Seeget_df_collection(), which returns an actually usableDFCollectioninstance (a subclass).-
MetaManager¶
-
class
dataCAT.context_managers.MetaManager(filename, manager)[source]¶ A wrapper for context managers.
Has a single important method,
MetaManager.open(), which calls and returns the context manager stored inMetaManager.manager.Note
MetaManager.filenamewill be the first positional argument provided toMetaManager.manager.Parameters: - filename (str) – The path+filename of a database component
See
MetaManager.filename. - manager (type [AbstractContextManager]) – A type object of a context manager.
TThe first positional argument of the context manager should be the filename.
See
MetaManager.manager.
-
manager¶ A type object of a context manager. The first positional argument of the context manager should be the filename.
Type: type [AbstractContextManager]
-
open(*args, **kwargs)[source]¶ Call and return
MetaManager.manager.Parameters: - *args – Positional arguments for
MetaManager.manager. - **kwargs – Keyword arguments for
MetaManager.manager.
Returns: An instance of a context manager.
Return type: AbstractContextManager_
- *args – Positional arguments for
- filename (str) – The path+filename of a database component
See
OpenLig¶
-
class
dataCAT.context_managers.OpenLig(filename=None, write=True)[source]¶ Context manager for opening and closing the ligand database (
Database.csv_lig).Parameters: -
df¶ An attribute for (temporary) storing the opened .csv file (see
OpenLig.filename) as aDataFrameinstance.Type: None or pd.DataFrame
-
OpenQD¶
-
class
dataCAT.context_managers.OpenQD(filename=None, write=True)[source]¶ Context manager for opening and closing the QD database (
Database.csv_qd).Parameters: -
df¶ An attribute for (temporary) storing the opened .csv file (
OpenQD.filename) asDataFrameinstance.Type: None or pd.DataFrame
-
OpenYaml¶
-
class
dataCAT.context_managers.OpenYaml(filename=None, write=True)[source]¶ Context manager for opening and closing job settings (
Database.yaml).Parameters: -
settings¶ An attribute for (temporary) storing the opened .yaml file (
OpenYaml.filename) asSettingsinstance.Type: None or plams.Settings
-
Function API¶
-
dataCAT.df_collection.get_df_collection(df)[source]¶ Return a mutable collection for holding dataframes.
Parameters: df (pd.DataFrame) – A Pandas DataFrame. Returns: A DFCollectioninstance. The class is described in more detail in the documentation of its superclass:_DFCollection.Return type: dataCAT.DFCollection_ Note
As the
DFCollectionclass is defined within the scope of this function, two instances ofDFCollectionwill not belong to the same class (see example below). In more technical terms: The class bound to a particularDFCollectioninstance is a unique instance oftype.>>> import numpy as np >>> import pandas as pd >>> df = pd.DataFrame(np.random.rand(5, 5)) >>> collection1 = get_df_collection(df) >>> collection2 = get_df_collection(df) >>> print(df is collection1.df is collection2.df) True >>> print(collection1.__class__.__name__ == collection2.__class__.__name__) True >>> print(collection1.__class__ == collection2.__class__) False
-
dataCAT.database_functions.as_pdb_array(mol_list, min_size=0)[source]¶ Convert a list of PLAMS molecule into an array of (partially) de-serialized .pdb files.
Parameters: - mol_list (\(m\) list [plams.Molecule]) – A list of \(m\) PLAMS molecules.
- min_size (int) – The minimumum length of the pdb_array. The array is padded with empty strings if required.
Returns: An array with \(m\) partially deserialized .pdb files with up to \(n\) lines each.
Return type: \(m*n\) np.ndarray [np.bytes |S80]
-
dataCAT.database_functions.from_pdb_array(array, rdmol=True)[source]¶ Convert an array with a (partially) de-serialized .pdb file into a molecule.
Parameters: - array (\(n\) np.ndarray [np.bytes / S80]) – A (partially) de-serialized .pdb file with \(n\) lines.
- rdmol (bool) – If
True, return an RDKit molecule instead of a PLAMS molecule.
Returns: A PLAMS or RDKit molecule build from array.
Return type: plams.Molecule or rdkit.Chem.Mol_
-
dataCAT.database_functions.sanitize_yaml_settings(settings, job_type)[source]¶ Remove a predetermined set of unwanted keys and values from a settings object.
Parameters: - settings (plams.Settings) – A settings instance with, potentially, undesired keys and values.
- job_type (str) – The name of key in the settings blacklist.
Returns: A new Settings instance with all unwanted keys and values removed.
Return type: plams.Settings_
Raises: KeyError – Raised if jobtype is not found in …/CAT/data/templates/settings_blacklist.yaml.