Bond Dissociation Energy¶
Calculate the bond dissociation energy (BDE) of ligands attached to the surface of the core. The calculation consists of five distinct steps:
1. Dissociate all combinations of \({n}\) ligands (\(Y\), see
optional.qd.dissociate.lig_count) a nd an atom from the core (\(X\), seeoptional.qd.dissociate.core_atom) within a radius \(r\) from aforementioned core atom (seeoptional.qd.dissociate.lig_core_distandoptional.qd.dissociate.core_core_dist). The dissociated compound has the general structure of \(XY_{n}\).2. Optimize the geometry of \(XY_{n}\) at the first level of theory (\(1\)). Default: ADF MOPAC [1, 2, 3].
3. Calculate the “electronic” contribution to the BDE (\(\Delta E\)) at the first level of theory (\(1\)): ADF MOPAC [1, 2, 3]. This step consists of single point calculations of the complete quantum dot, \(XY_{n}\) and all \(XY_{n}\)-dissociated quantum dots.
4. Calculate the thermalchemical contribution to the BDE (\(\Delta \Delta G\)) at the second level of theory (\(2\)). Default: ADF UFF [4, 5]. This step consists of geometry optimizations and frequency analyses of the same compounds used for step 3.
- \(\Delta G_{tot} = \Delta E_{1} + \Delta \Delta G_{2} = \Delta E_{1} + (\Delta G_{2} - \Delta E_{2})\).
Default Settings¶
optional:
qd:
dissociate:
core_atom: Cd
lig_count: 2
keep_files: True
core_core_dist: 5.0 # Ångström
lig_core_dist: 5.0 # Ångström
core_index: False
topology: {}
job1: AMSJob
s1: True
job2: AMSJob
s2: True
Arguments¶
-
optional.qd.dissociate optional: qd: dissociate: core_atom: Cd lig_count: 2 keep_files: True core_core_dist: 5.0 # Ångström lig_core_dist: 5.0 # Ångström core_index: False topology: 7: vertice 8: edge 10: face
optional.qd.dissociate.core_atom¶
Parameter: The atomic number or atomic symbol of the core atoms (\(X\)) which are to be dissociated. The core atoms are dissociated in combination with \(n\) ligands (\(Y\), see
optional.qd.dissociate.lig_count). Yields a compound with the general formula \(XY_{n}\).If one is interested in dissociating ligands in combination with a molecular species (e.g. \(X = {NR_4}^+\)) the atomic number (or symbol) can be substituted for a SMILES string represting a poly-atomic ion (e.g. tetramethyl ammonium: C[N+](C)(C)C).
If a SMILES string is provided it must satisfy the following 2 requirements:
- The SMILES string must contain a single charged atom; unpredictable behaviour can occur otherwise.
- The provided structure (including its bonds) must be present in the core.
Warning
This argument has no value be default and thus must be provided by the user.
Note
The yaml format uses
nullrather thanNoneas in Python.
optional.qd.dissociate.lig_count¶
Parameter:
- Type -
int- Default value –
NoneThe number of ligands, \(n\), which is to be dissociated in combination with a single core atom (\(X\), see
optional.qd.dissociate.core_atom). Yields a compound with the general formula \(XY_{n}\).Warning
This argument has no value be default and thus must be provided by the user.
Note
The yaml format uses
nullrather thanNoneas in Python.
optional.qd.dissociate.keep_files¶
Parameter:
- Type -
bool- Default value –
TrueWhether to keep or delete all BDE files after all calculations are finished.
optional.qd.dissociate.core_core_dist¶
Parameter: The maximum to be considered distance (Ångström) between atoms in
optional.qd.dissociate.core_atom. Used for determining the topology of the core atom (seeoptional.qd.dissociate.topology) and whether it is exposed to the surface of the core or not. It is recommended to use a radius which encapsulates a single (complete) shell of neighbours.If not specified (or equal to
0.0) CAT will attempt to guess a suitable value based on the cores’ radial distribution function.
optional.qd.dissociate.lig_core_dist¶
Parameter: Dissociate all possible combinations of \(n\) ligands and a single core atom (see
optional.qd.dissociate.core_atom) within a given radius (Ångström) from aforementioned core atom. The number of ligands dissociated in combination with a single core atom is controlled byoptional.qd.dissociate.lig_count.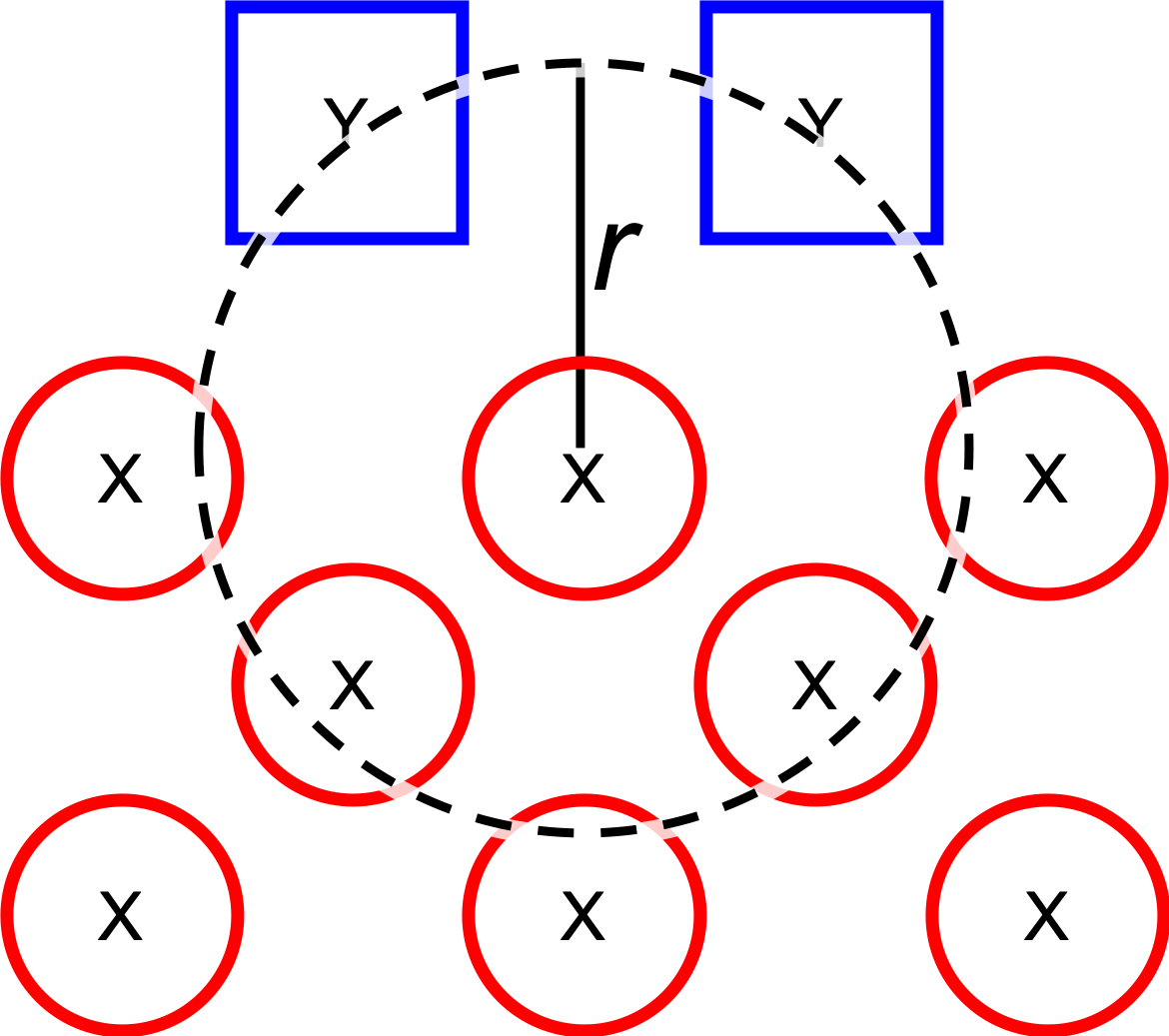
optional.qd.dissociate.core_index¶
Parameter: Alternative to
optional.qd.dissociate.lig_core_distandoptional.qd.dissociate.core_atom. Manually specify the indices of all to-be dissociated atoms in the core. Core atoms will be dissociated in combination with the \(n\) closest ligands.Note
The yaml format uses
nullrather thanNoneas in Python.
optional.qd.dissociate.topology¶
Parameter:
- Type -
dict- Default value –
{}A dictionary which translates the number neighbouring core atoms (see
optional.qd.dissociate.core_atomandoptional.qd.dissociate.core_core_dist) into a topology. Keys represent the number of neighbours, values represent the matching topology.Example
Given a
optional.qd.dissociate.core_core_distof5.0Ångström, the following options can be interpreted as following:optional: qd: dissociate: 7: vertice 8: edge 10: faceCore atoms with
7other neighbouring core atoms (within a radius of5.0Ångström) are marked as"vertice", the ones with8neighbours are marked as"edge"and the ones with10neighbours as"face".
Arguments - Job Customization¶
-
optional.qd.dissociate optional: qd: dissociate: job1: AMSJob s1: True job2: AMSJob s2: True
optional.qd.dissociate.job1¶
Parameter: A
typeobject of aplams.Jobsubclass, used for calculating the “electronic” component (\(\Delta E_{1}\)) of the bond dissociation energy. Involves single point calculations.Alternatively, an alias can be provided for a specific job type (see Type Aliases).
Setting it to
Truewill default toplams.AMSJob, whileFalseis equivalent tooptional.qd.dissociate=False.
optional.qd.dissociate.s1¶
Parameter: The job settings used for calculating the “electronic” component (\(\Delta E_{1}\)) of the bond dissociation energy.
Alternatively, a path can be provided to .json or .yaml file containing the job settings.
Setting it to
Truewill default to the["MOPAC"]block in CAT/data/templates/qd.yaml, whileFalseis equivalent tooptional.qd.dissociate=False.
optional.qd.dissociate.job2¶
Parameter: A
typeobject of aplams.Jobsubclass, used for calculating the thermal component (\(\Delta \Delta G_{2}\)) of the bond dissociation energy. Involves a geometry reoptimizations and frequency analyses.Alternatively, an alias can be provided for a specific job type (see Type Aliases).
Setting it to
Truewill default toplams.AMSJob, whileFalsewill skip the thermochemical analysis completely.
optional.qd.dissociate.s1
Parameter: The job settings used for calculating the thermal component (\(\Delta \Delta G_{2}\)) of the bond dissociation energy.
Alternatively, a path can be provided to .json or .yaml file containing the job settings.
Setting it to
Truewill default to the the MOPAC block in CAT/data/templates/qd.yaml, whileFalsewill skip the thermochemical analysis completely.