The Database Class
A Class designed for the storing, retrieval and updating of results.
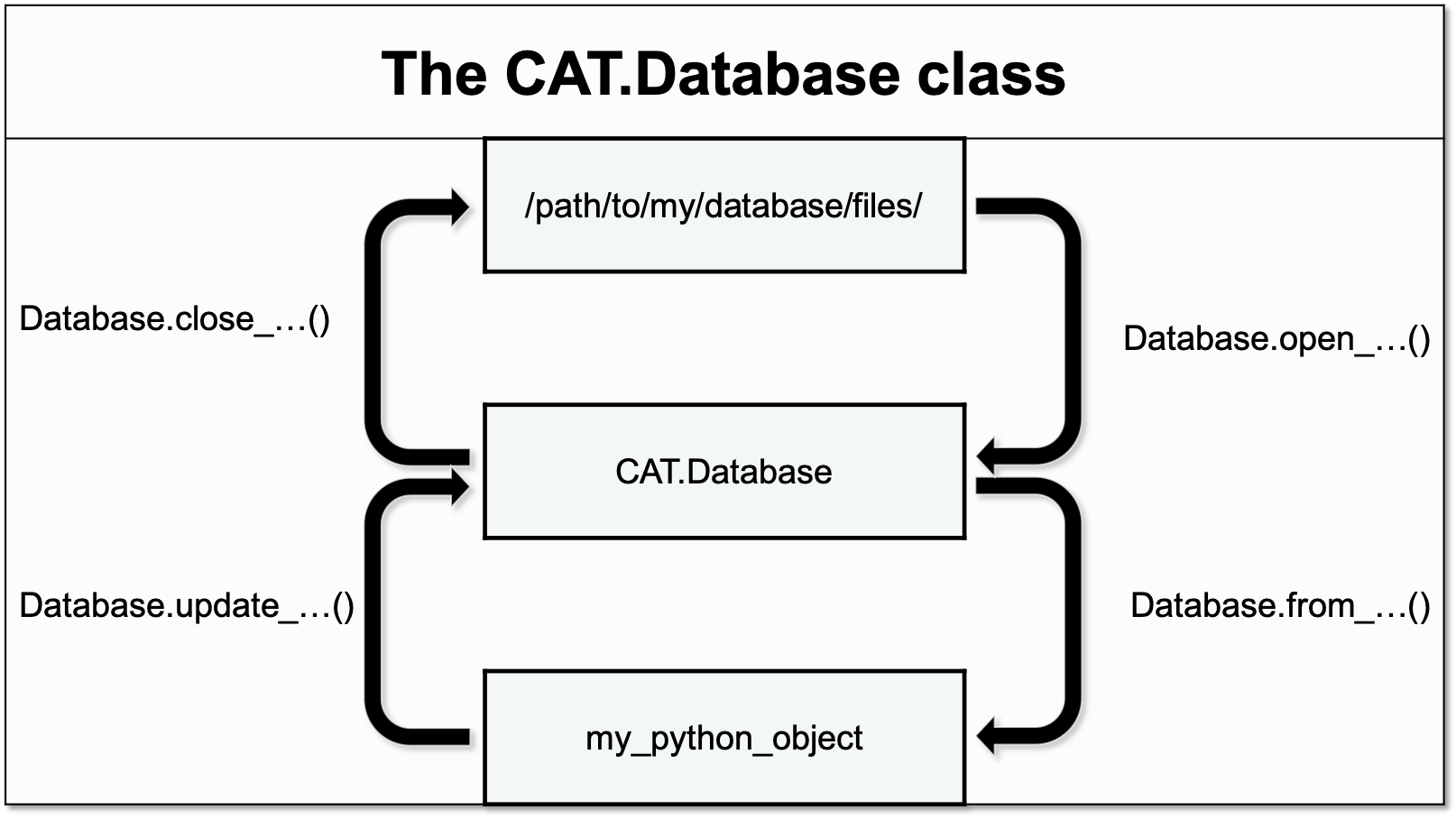
The methods of the Database class can be divided into three categories accoring to their functionality:
Opening & closing the database - these methods serve as context managers for loading and unloading parts of the database from the harddrive.
The context managers can be accessed by calling either
Database.csv_lig,Database.csv_qd, orDatabase.hdf5, with the option of passing additional positional or keyword arguments.>>> from dataCAT import Database >>> database = Database() >>> with database.csv_lig(write=False) as db: >>> print(repr(db)) DFProxy(ndframe=<pandas.core.frame.DataFrame at 0x7ff8e958ce80>) >>> with database.hdf5('r') as db: >>> print(type(db)) <class 'h5py._hl.files.File'>
Importing to the database - these methods handle the importing of new data from python objects to the Database class:
Exporting from the database - these methods handle the exporting of data from the Database class to other python objects or remote locations:
Index
Get the path+filename of the directory containing all database components. |
|
Get a function for constructing an |
|
Get a function for constructing an |
|
Get a function for constructing a |
|
Get a mapping with keyword arguments for |
|
|
Export ligand or qd results to the MongoDB database. |
|
Update |
|
Export molecules (see the |
|
Pull results from |
|
Import structures from the hdf5 database as RDKit or PLAMS molecules. |
|
A mutable wrapper providing a view of the underlying dataframes. |
|
Context manager for opening and closing the ligand database ( |
|
Context manager for opening and closing the QD database ( |
API
- class dataCAT.Database(path=None, host='localhost', port=27017, **kwargs)[source]
The Database class.
- property dirname
Get the path+filename of the directory containing all database components.
- property csv_lig
Get a function for constructing an
dataCAT.OpenLigcontext manager.
- property csv_qd
Get a function for constructing an
dataCAT.OpenQDcontext manager.
- property mongodb
Get a mapping with keyword arguments for
pymongo.MongoClient.- Type
Mapping[str, Any], optional
- update_mongodb(database='ligand', overwrite=False)[source]
Export ligand or qd results to the MongoDB database.
Examples
>>> from dataCAT import Database >>> kwargs = dict(...) >>> db = Database(**kwargs) # Update from db.csv_lig >>> db.update_mongodb('ligand') # Update from a lig_df, a user-provided DataFrame >>> db.update_mongodb({'ligand': lig_df}) >>> print(type(lig_df)) <class 'pandas.core.frame.DataFrame'>
- Parameters
database (
strorMapping[str, pandas.DataFrame]) – The type of database. Accepted values are"ligand"and"qd", openingDatabase.csv_ligandDatabase.csv_qd, respectivelly. Alternativelly, a dictionary with the database name and a matching DataFrame can be passed directly.overwrite (
bool) – Whether or not previous entries can be overwritten or not.
- Return type
- update_csv(df, index=None, database='ligand', columns=None, overwrite=False, job_recipe=None, status=None)[source]
Update
Database.csv_ligorDatabase.csv_qdwith new settings.- Parameters
df (
pandas.DataFrame) – A dataframe of new (potential) database entries.database (
str) – The type of database; accepted values are"ligand"(Database.csv_lig) and"qd"(Database.csv_qd).columns (
Sequence, optional) – Optional: A sequence of column keys in df which (potentially) are to be added to this instance. IfNoneAdd all columns.overwrite (
bool) – Whether or not previous entries can be overwritten or not.status (
str, optional) – A descriptor of the status of the moleculair structures. Set to"optimized"to treat them as optimized geometries.
- Return type
- update_hdf5(df, index, database='ligand', overwrite=False, status=None)[source]
Export molecules (see the
"mol"column in df) to the structure database.Returns a series with the
Database.hdf5indices of all new entries.- Parameters
df (
pandas.DataFrame) – A dataframe of new (potential) database entries.database (
str) – The type of database; accepted values are"ligand"and"qd".overwrite (
bool) – Whether or not previous entries can be overwritten or not.status (
str, optional) – A descriptor of the status of the moleculair structures. Set to"optimized"to treat them as optimized geometries.
- Returns
A series with the indices of all new molecules in
Database.hdf5.- Return type
- from_csv(df, database='ligand', get_mol=True, inplace=True)[source]
Pull results from
Database.csv_ligorDatabase.csv_qd.Performs in inplace update of df if inplace =
True, thus returingNone.- Parameters
df (
pandas.DataFrame) – A dataframe of new (potential) database entries.database (
str) – The type of database; accepted values are"ligand"and"qd".get_mol (
bool) – Attempt to pull preexisting molecules from the database. See the inplace argument for more details.inplace (
bool) – IfTrueperform an inplace update of the"mol"column in df. Otherwise return a new series of PLAMS molecules.
- Returns
Optional: A Series of PLAMS molecules if get_mol =
Trueand inplace =False.- Return type
pandas.Series, optional
- from_hdf5(index, database='ligand', rdmol=True, mol_list=None)[source]
Import structures from the hdf5 database as RDKit or PLAMS molecules.
- Parameters
index (
Sequence[int]orslice) – The indices of the to be retrieved structures.database (
str) – The type of database; accepted values are"ligand"and"qd".rdmol (
bool) – IfTrue, return an RDKit molecule instead of a PLAMS molecule.
- Returns
A list of PLAMS or RDKit molecules.
- Return type
- hdf5_availability(timeout=5.0, max_attempts=10)[source]
Check if a .hdf5 file is opened by another process; return once it is not.
If two processes attempt to simultaneously open a single hdf5 file then h5py will raise an
OSError.The purpose of this method is ensure that a .hdf5 file is actually closed, thus allowing the
Database.from_hdf5()method to safely access filename without the risk of raising anOSError.- Parameters
timeout (
float) – Time timeout, in seconds, between subsequent attempts of opening filename.max_attempts (
int, optional) – Optional: The maximum number attempts for opening filename. If the maximum number of attempts is exceeded, raise anOSError. Setting this value toNonewill set the number of attempts to unlimited.
- Raises
OSError – Raised if max_attempts is exceded.
See also
dataCAT.functions.hdf5_availability()This method as a function.
- class dataCAT.DFProxy(ndframe)[source]
A mutable wrapper providing a view of the underlying dataframes.
- ndframe
The embedded DataFrame.
- Type
- class dataCAT.OpenLig(filename, write=True)[source]
Context manager for opening and closing the ligand database (
Database.csv_lig).
- class dataCAT.OpenQD(filename, write=True)[source]
Context manager for opening and closing the QD database (
Database.csv_qd).